Login functionality is undergoing maintenance. If you encounter problems while ordering, please contact us at BDBCustomerService@bd.com.
Sunday, May 24, 2026, 5:30 - 8:30 pm (HKT)
See More
Login functionality is undergoing maintenance. If you encounter problems while ordering, please contact us at BDBCustomerService@bd.com.
Sunday, May 24, 2026, 5:30 - 8:30 pm (HKT)
See MoreLogin functionality is undergoing maintenance. If you encounter problems while ordering, please contact us at BDBCustomerService@bd.com.
Sunday, May 24, 2026, 5:30 - 8:30 pm (HKT)
.Looks like you're visiting us from {countryName}.
Would you like to stay on the current location or be switched to your location?
The BD Rhapsody™ Sequence Analysis Pipeline is a versatile tool that offers the flexibility to run your bioinformatics analysis on either a Seven Bridges cloud-based platform or on a local installation.
The BD Rhapsody™ Sequence Analysis Pipeline:
The .Cellismo output files from the BD Rhapsody™ Sequence Analysis Pipeline can be imported into the BD Cellismo™ Data Visualization Tool for secondary analysis and visualization.
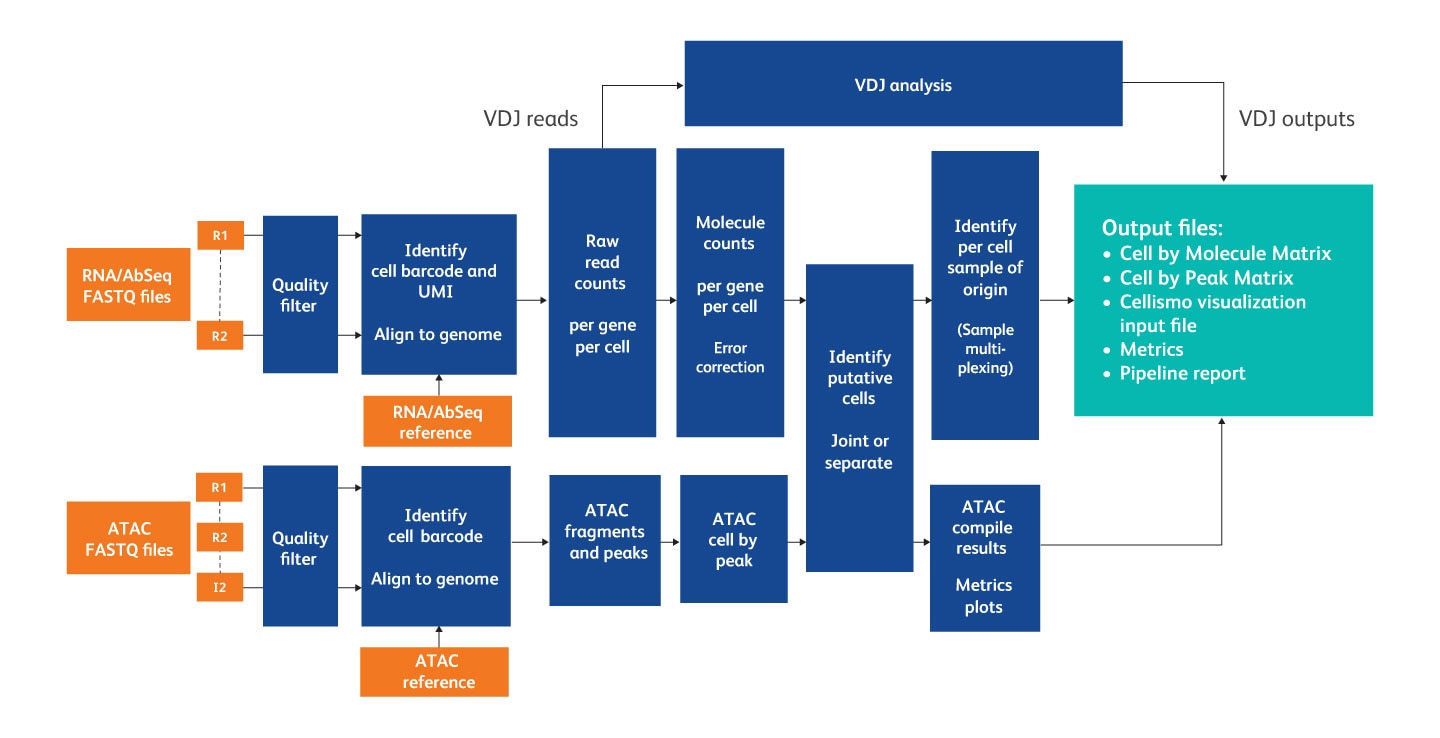
After sequencing, the pipeline takes input from FASTQ files, a reference (Targeted panel or WTA / WTA+ATAC-Seq reference archive), an AbSeq reference (if required) and a supplemental reference (if required) to generate output files and metrics about the pipeline run.
Overview of the steps in the analysis pipeline.
v3 BD Rhapsody™ Sequence Analysis Pipeline | October 2025
Added
Updated
Fixed
Cloud-Based Version
Local Version
For Research Use Only. Not for use in diagnostic or therapeutic procedures.