-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD OMICS-Guard™ CRYO Preservation Buffer
- BD OMICS-Guard™ Sample Preservation Buffer
- BD OMICS-One™ Protein Panels
- BD® AbSeq Assay
- BD® Single-Cell Multiplexing Kit
- BD OMICS-One™ WTA Next Assay
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Accessory Kits
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
-
Training
- Flow Cytometry Basic Training
-
Product-Based Training
- BD Accuri™ C6 Plus Cell Analyzer
- BD FACSAria™ Cell Sorter Cell Sorter
- BD FACSCanto™ Cell Analyzer
- BD FACSDiscover™ A8 Cell Analyzer
- BD FACSDiscover™ S8 Cell Sorter
- BD FACSDuet™ Sample Preparation System
- BD FACSLyric™ Cell Analyzer
- BD FACSMelody™ Cell Sorter
- BD FACSymphony™ Cell Analyzer
- BD LSRFortessa™ Cell Analyzer
- Advanced Training
Old Browser
This page has been recently translated and is available in French now.
Looks like you're visiting us from {countryName}.
Would you like to stay on the current location site or be switched to your location?
Expanding the Capabilities of Single-cell Experiments Using Customizable Beads
Conventional single-cell workflows rely almost exclusively on the capture of poly(A)-tailed mRNA, which leads to the possibility of missing target sequences. The ability to modify the beads to target sequences that do not rely on poly-adenylation allows for simultaneous profiling of standard transcriptomes and specific regions of interest that can expand the capabilities of single cell experiments.1
In a 2025 study published in Nature Communications (Moro et al. 2025),2 researchers modified BD Rhapsody™ Enhanced Beads to detect BCR::ABL1 fusion transcripts, a primary driver in acute lymphoblastic leukemia. A mix of three patient-derived leukemia cell lines were used that either harbored a major fusion breakpoint in BCR (e14a2 in K562 cells), a minor fusion breakpoint (e1a2 in SUP-B15 cells), or no fusion transcript (Loucy cells). Using the published data in combination with the BD Cellismo™ Data Visualization Tool, figures were generated to illustrate the detection of these fusion transcripts in the corresponding cell lines.
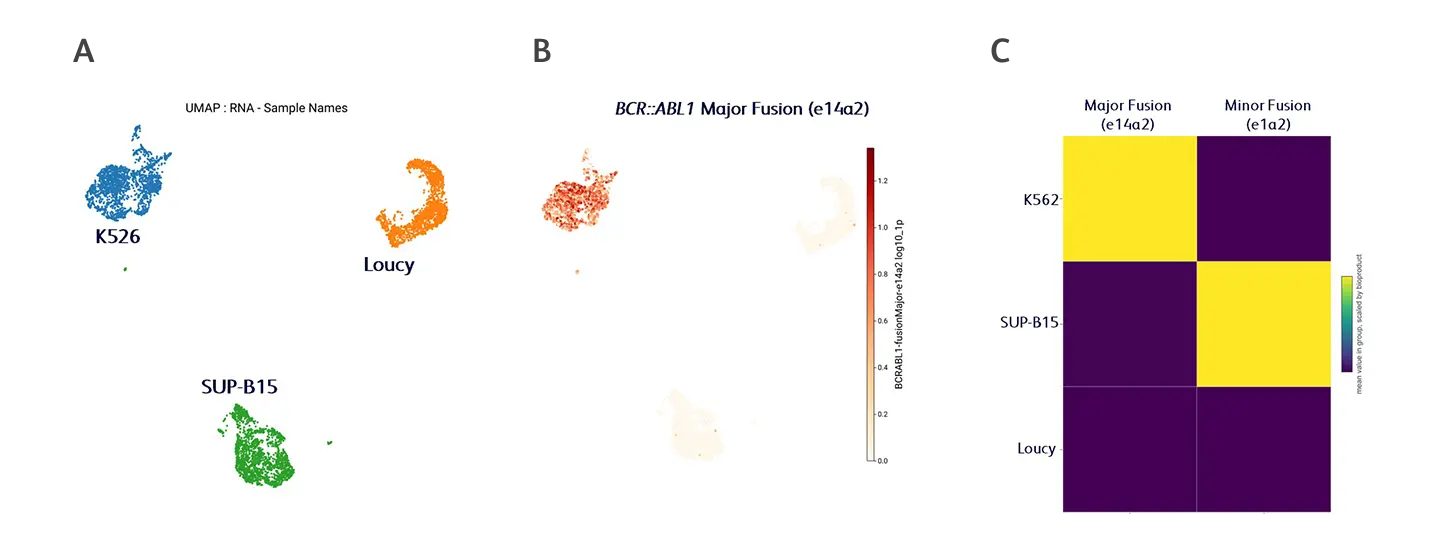
Detection and profiling of fusion transcripts through internal transcript capture. (A) UMAP projection driven by whole‑transcriptome mRNA expression showing clear separation of the cell lines. (B) BCR::ABL1 major fusion transcript (e14a2) expression in the corresponding K526 cell line, as expected. (C) Heatmap illustrating the distribution of both major and minor BCR::ABL1 fusion transcripts across all cell lines analyzed.
Expanding the Scope of single-cell analysis using the BD Rhapsody™ Enhanced Beads
The BD Rhapsody™ Enhanced Beads can be easily modified by a user to target unique sequences. By leveraging modified beads, a variety of non-canonical targets can be uniquely captured including sequences that lack poly(A) tails:
- Non-polyadenylated RNA: lncRNAs, snRNAs/snoRNA, miRNAs etc.
- Internal transcript sequences: specific regions associated with gene fusions or specific sequence variants.
- Guide RNAs: essential for single-cell CRISPR screening applications.
- Viral RNA: including viral genomes, sub-genomic RNAs, or specific viral transcripts.
- Low-abundance RNA: capture of rare transcripts like transcription factors or signaling molecules.
Innovative Bead Architecture
Unlike gel-based or dissolvable beads, the BD Rhapsody™ Enhanced Beads contain a solid-core and are magnetic, allowing for precise chemical and thermal manipulation. Each bead features a dual-oligonucleotide design:
- Poly(dT) oligonucleotide: captures 3' poly(A) tails of mRNA transcripts for standard whole-transcriptome (WTA) or targeted analysis, as well as oligonucleotide-tagged antibodies (CITE-Seq and cell-hashing).
- Customizable Template Switch Oligonucleotide (TSO): harbors a unique 3’-end sequence that can be modified to incorporate custom capture sequences, enabling selective enrichment of noncanonical or user-defined targets while still preserving the ability to perform standard transcriptome analysis in parallel. ‑canonical or user‑defined targets while still preserving the ability to perform standard transcriptome analysis in parallel.
Because these two capture sites are distinct, users can selectively enrich custom targets and prepare them as independent libraries without compromising the integrity of the standard mRNA library.
Key Bead Features and Workflow
- Robust solid structure: Beads are magnetic, enabling rapid and efficient separation during buffer exchanges and wash steps, simplifying overall handling.
- Heat-stable: Beads are heat stable (across 4 °C to 95 °C), and compatible with standard enzymatic reaction conditions, supporting a wide range of molecular workflows.
- Chemical compatibility: Beads tolerate a range of chemical modifications including NaOH treatment, allowing for additional modification approaches.
- Longterm stability: Beads are stable for at least one year at 4 °C after cDNA synthesis. They can be subsampled prior to processing, enabling reduced sequencing and lower overall experimental costs.
Simple Workflow
The schematic below highlights the minimal steps required to customize the BD Rhapsody™ Enhanced Beads for unique target capture.
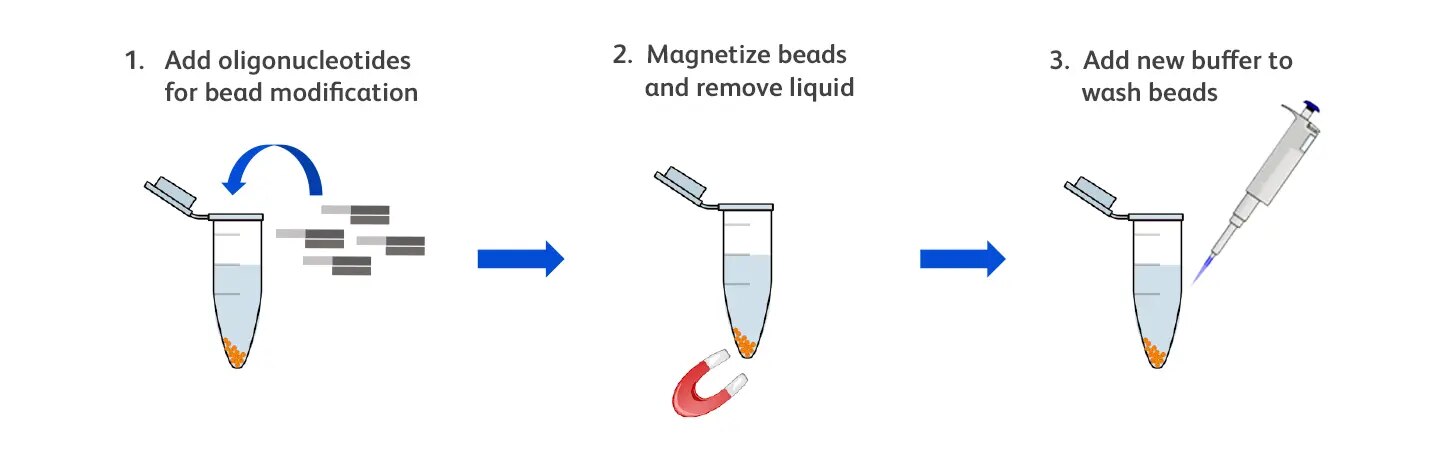
Bead modification can involve three simple steps: (1) adding custom oligonucleotides to the beads and incubating to allow annealing, (2) magnetizing the beads and removing the supernatant, and (3) washing the beads by adding fresh buffer.
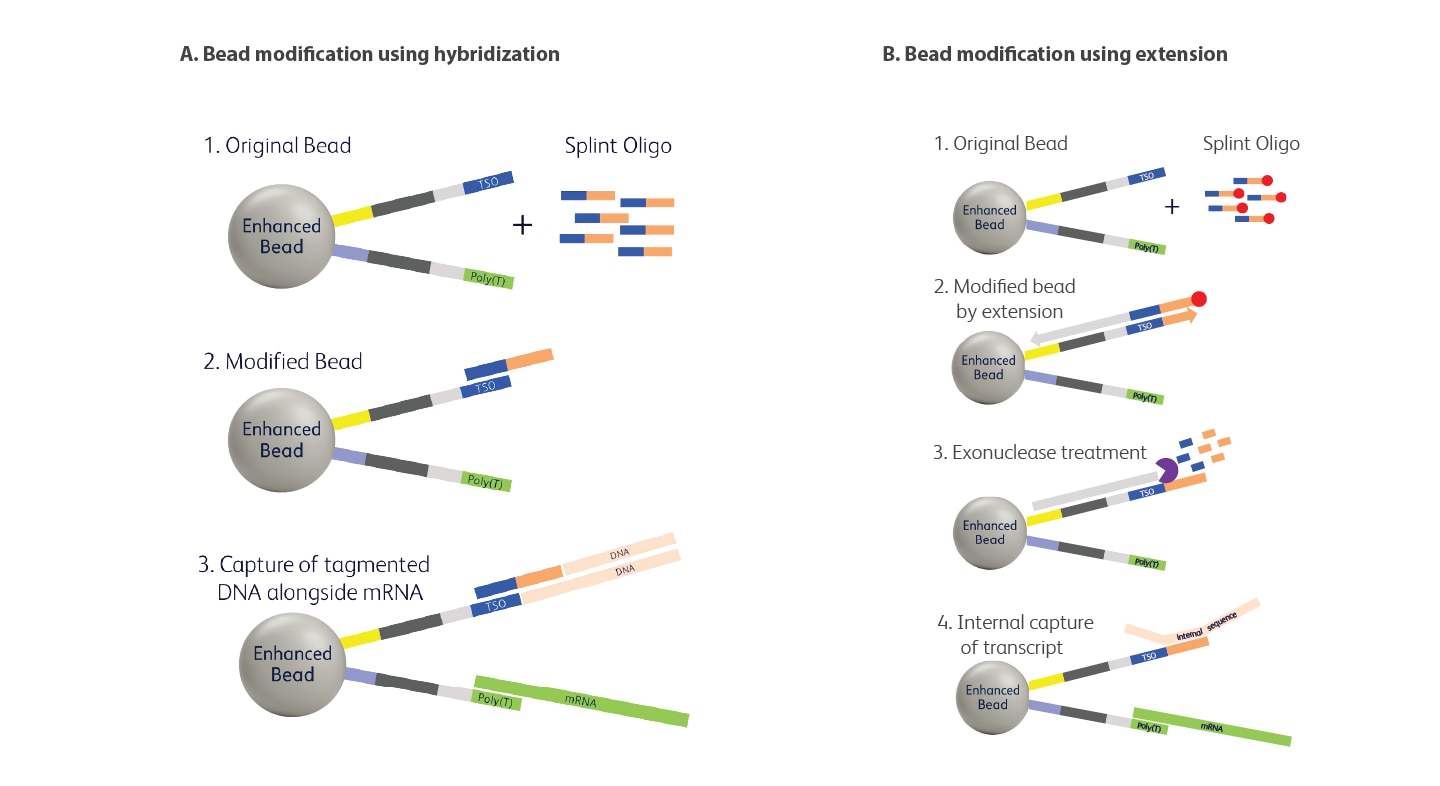
Two approaches for modifying the bead. (A) The splint oligo is annealed to the bead bound capture sequence, where it remains hybridized during capture of the desired target molecules. This strategy is used in the BD Rhapsody™ ATAC-seq assays (see BD Rhapsody single-cell ATAC-seq protocols) to isolate tagmented DNA fragments. A recent publication in Cell Reports (Kohne et al., 2025) illustrates the use of this type of modification. (B) The bead is modified through enzymatic extension, using a splint oligo as a template to generate a new capture sequence on the bead. This approach was used in the Moro et. al., 2025 Nature Communications study. Note: red circle indicates a 5’ phosphate group; in the published method, an additional oligonucleotide was included to protect the poly(dT) strand; not shown here.
Potential Applications
The examples shown above represent only a subset of the applications enabled by bead modification. Additional possibilities may include:
- Organelle and mitochondrial transcript capture
- Immunology and CRISPR perturbation screens
- Host-pathogen dual transcriptomics
- Developmental biology and stem cell transcript profiling
- Microbiome and environmental single‑cell studies
These examples highlight how the ability to engineer BD Rhapsody™ enhanced Beads for orthogonal capture of non-canonical and sequence-specific targets substantially broadens the analytical resolution of single-cell workflows, thus enabling deeper molecular profiling.
Read the full Nature Communications paper about the RoCK and ROI approach.
References
- Dincaslan FB, Ngang SWY, Tan RZ and Cheow LF. Automated high-throughput profiling of single-cell total transcriptome with scComplete-seq. 2025; Nucleic Acids Research, 53:14, gkaf699. Doi: 10.1093/nar/gkaf699
- Moro G, Mallona I, Barz MJ, Maillard J, Brugger MD, et al. RoCK and ROI: single-cell transcriptomics with multiplexed enrichment of selected transcripts and region-specific sequencing. 2025. Nature Comm; 16:10991. https://doi.org/10.1038/s41467-025-66248-z
For Research Use Only. Not for use in diagnostic or therapeutic procedures.