-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD® AbSeq Assay
- BD Rhapsody™ Accessory Kits
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Profiling Assays
- BD OMICS-Guard Sample Preservation Buffer
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
-
- BD® AbSeq Assay
- BD Rhapsody™ Accessory Kits
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Profiling Assays
- BD OMICS-Guard Sample Preservation Buffer
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- Korea (Korea)
-
국가 / 언어 변경
Old Browser
Looks like you're visiting us from {countryName}.
Would you like to stay on the current country site or be switched to your country?
![FITC Rat Anti-Mouse Vα 3.2[b,c] TCR](/content/dam/bdb/products/global/reagents/flow-cytometry-reagents/research-reagents/single-color-antibodies-ruo/553xxx/5532xx/553219_base/01454C_553219_image1.png)
![FITC Rat Anti-Mouse Vα 3.2[b,c] TCR](/content/dam/bdb/products/global/reagents/flow-cytometry-reagents/research-reagents/single-color-antibodies-ruo/553xxx/5532xx/553219_base/01454C_553219_image1.png)
.png)
![FITC Rat Anti-Mouse Vα 3.2[b,c] TCR](/content/dam/bdb/products/global/reagents/flow-cytometry-reagents/research-reagents/single-color-antibodies-ruo/553xxx/5532xx/553219_base/01454C_553219_image1.png)
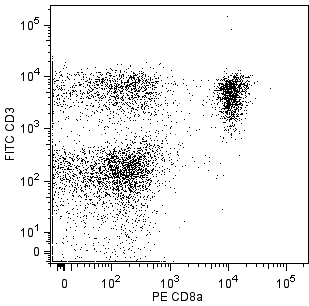
Two-color analysis of the expression of Vα 3.2b,c TCR on peripheral lymphocytes. C57BL/6 lymph node leukocytes were simultaneuosly stained with FITC-conjugated RR3-16, PE-conjugated anti-mouse CD4 RM4-5 (Cat. No. 553048/553049), and PE-conjugated anti-mouse CD8a 53-6.7 (Cat. No. 553032/553033) monoclonal antibodies. Flow cytometry was performed on a BD FACScan™ flow cytometry system.
.png)
![FITC Rat Anti-Mouse Vα 3.2[b,c] TCR](/content/dam/bdb/products/global/reagents/flow-cytometry-reagents/research-reagents/single-color-antibodies-ruo/553xxx/5532xx/553219_base/01454C_553219_image1.png)
BD Pharmingen™ FITC Rat Anti-Mouse Vα 3.2[b,c] TCR
.png)
규제 상태 범례
Becton, Dickinson and Company의 명시적인 서면 승인 없이는 사용 하실 수 없습니다.
준비 및 보관
제품 고시
- Since applications vary, each investigator should titrate the reagent to obtain optimal results.
- Please refer to www.bdbiosciences.com/us/s/resources for technical protocols.
- Caution: Sodium azide yields highly toxic hydrazoic acid under acidic conditions. Dilute azide compounds in running water before discarding to avoid accumulation of potentially explosive deposits in plumbing.
관련 제품
.png?imwidth=320)
.png?imwidth=320)
The RR3-16 monoclonal antibody specifically reacts with the Vα 3.2 T-Cell Receptor (TCR) of mice having the b (e.g., C57BL) and c (e.g., SWR, SJL, NZB, NOD) haplotypes of the Tcra gene complex, but not with TCR encoded by other members of Tcra-V3 gene subfamily. RR3-16 antibody does not react with strains having the a (e.g., A, AKR, BALB/c, CBA, C3H/He) or d (e.g., DBA/1, DBA/2, NZW) Tcra haplotypes. In addition, it has been shown that frequencies of Vα 3.2+ CD8+ T cells from homozygous H-2k/H-2k mice are moderately higher than those from heterozygous H-2k/H-2d mice, suggesting positive selection by H-2k.
This antibody is routinely tested by flow cytometric analysis. Other applications were tested at BD Biosciences Pharmingen during antibody development only or reported in the literature.
개발 참고 자료 (3)
-
Tomonari K, Fairchild S, Rosenwasser OA. Influence of viral superantigens on V beta- and V alpha-specific positive and negative selection. Immunol Rev. 1993; 131:131-168. (Biology). 참조 보기
-
Utsunomiya Y, Bill J, Palmer E, Gollob K, Takagaki Y, Kanagawa O. Analysis of a monoclonal rat antibody directed to the alpha-chain variable region (V alpha 3) of the mouse T cell antigen receptor. J Immunol. 1989; 143(8):2602-2608. (Immunogen). 참조 보기
-
Utsunomiya Y, Bill J, Palmer E, Kanagawa O. Identification of a mouse T-cell antigen receptor alpha-chain polymorphism by a V alpha 3.2 chain-specific monoclonal antibody. Immunogenetics. 1991; 33(3):198-201. (Clone-specific). 참조 보기
Please refer to Support Documents for Quality Certificates
Global - Refer to manufacturer's instructions for use and related User Manuals and Technical data sheets before using this products as described
Comparisons, where applicable, are made against older BD Technology, manual methods or are general performance claims. Comparisons are not made against non-BD technologies, unless otherwise noted.
For Research Use Only. Not for use in diagnostic or therapeutic procedures.
Report a Site Issue
This form is intended to help us improve our website experience. For other support, please visit our Contact Us page.