-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD OMICS-Guard™ Sample Preservation Buffer
- BD® AbSeq Assay
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Accessory Kits
- BD® OMICS-One Protein Panels
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
Old Browser
Looks like you're visiting us from {countryName}.
Would you like to stay on the current location site or be switched to your location?
For Professionals in Research
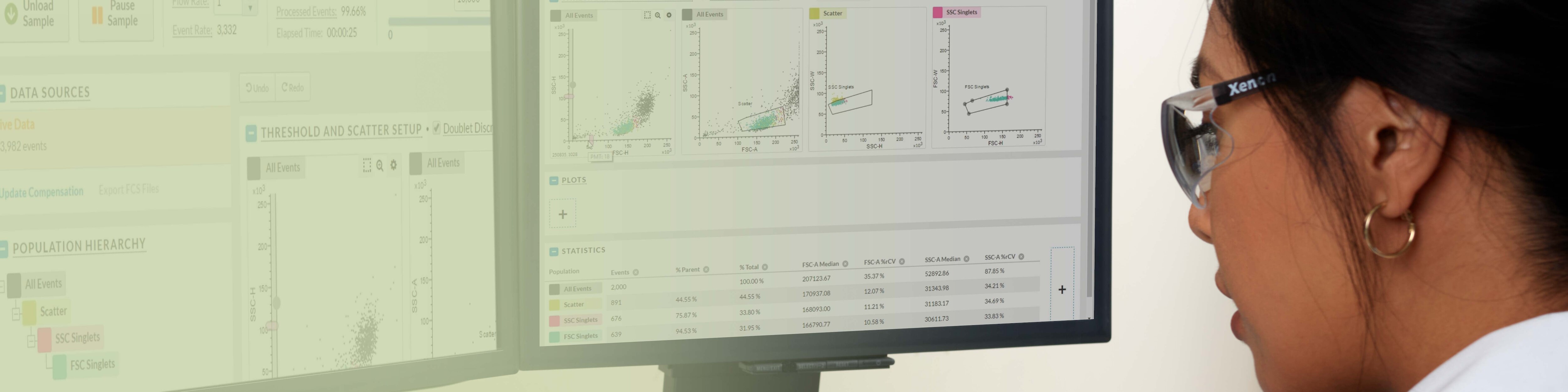
New Software for the Next Standard in Flow Cytometry
BD FACSChorus™ Software is designed to streamline and enhance your flow cytometry experience, making flow analysis and sorting more accessible and efficient as you grow to new heights with your BD FACSDiscover™ Platform instrument. With its intuitive interface and workflow-driven features, BD FACSChorus™ Software also gives flexibility and control where you need it, offering a comprehensive solution for instrument setup, quality control and data acquisition.
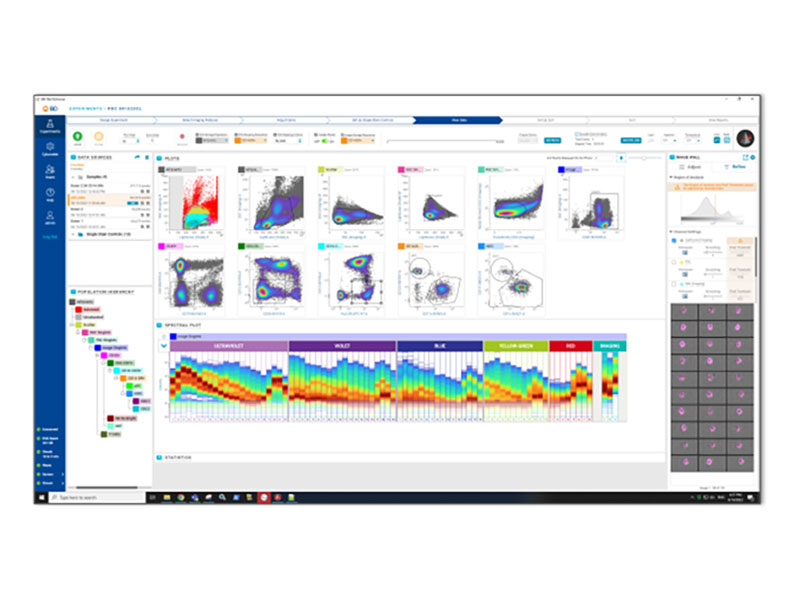
Enabling BD CellView™ Image Technology
and BD SpectralFX™ Technology on BD FACSDiscover™ Platform
BD FACSChorus™ Software brings BD SpectralFX™ Technology and BD CellView™ Image Technology to life for your samples.
BD FACSChorus™ Software brings an intuitive workflow and a high-powered system-aware unmixing algorithm to any spectral flow cytometry experiment. It simplifies the process of creating unmixing matrices and streamlines your experiments with flexible controls, extraction of multiple autofluorescent populations and post-acquisition reunmixing.
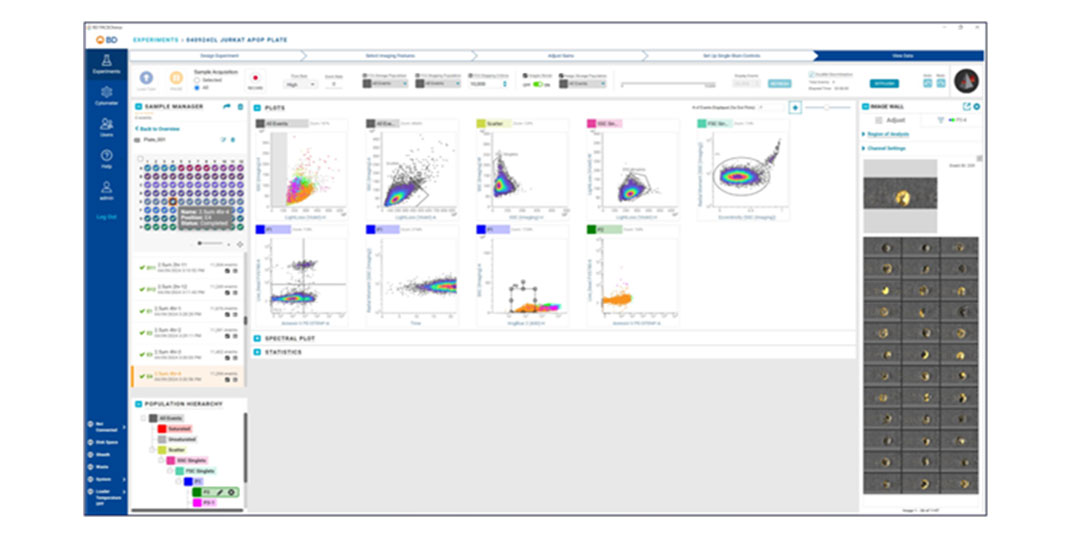
Applications
BD CellView™ Image Technology offers real-time visualization of cell images within BD FACSChorus™ Software. Users can mouse over cell events on plots or view them on the Image Wall in real time. This technology combines traditional flow cytometry parameters with imaging parameters to characterize populations, providing a comprehensive view of cell characteristics.
BD FACSChorus™ Software facilitates your analyzer and sorter workflows efficiently from startup to shutdown. With guided prompts for all setup, quality control, and maintenance procedures, it simplifies the operation of your BD FACSDiscover™ Platform. It provides a common workflow to your experiments, intuitive tool tips and instructions, ensuring an easy-to-use experience for all users. And when you’re ready to move your experiment from your analyzer to your sorter, simply transfer the experiment template and get ready to take your new cells to your downstream applications.
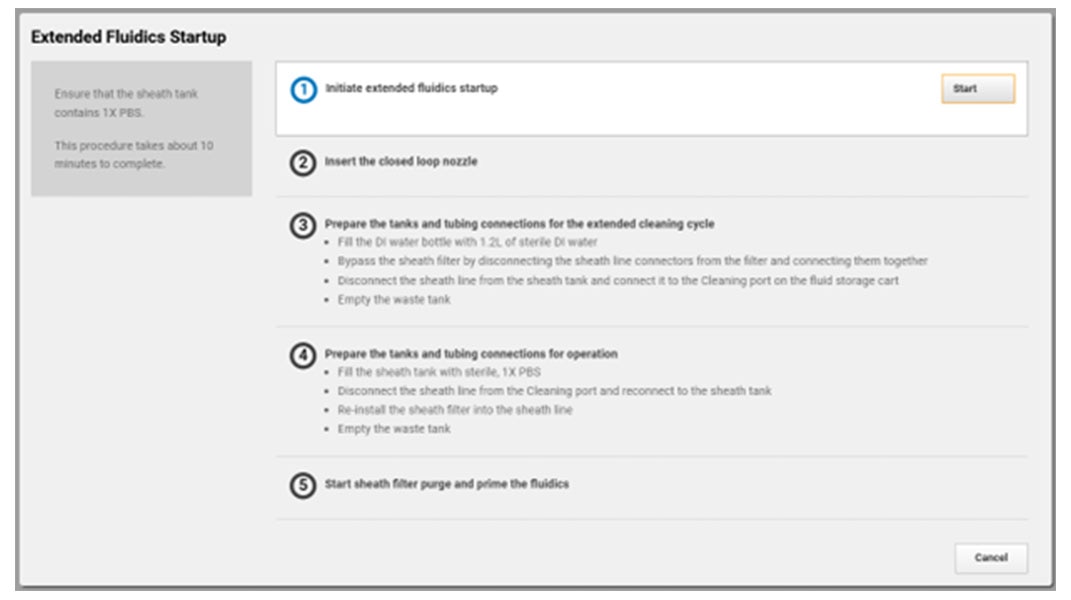
You’re busy trying to get the answers you need, BD FACSChorus™ Software helps manage both the BD FACSDiscover™ A8 Cell Analyzer and BD FACSDiscover™ S8 Cell Sorter with features for easier setup and use. On the sorter, the automated stream setup and side stream tracking simplifies sort setup, while features like adjusting the side stream targets and drop delay offset give you the flexibility you need.
On the analyzer, customize your plate or carrier-based acquisition with temperature and mixing settings before walking away while the acquisition occurs. The automated clog, empty detection and recovery with the BD FACSDiscover™ A8 Cell Analyzer will protect your samples and give you peace of mind in either imaging or high-speed mode.
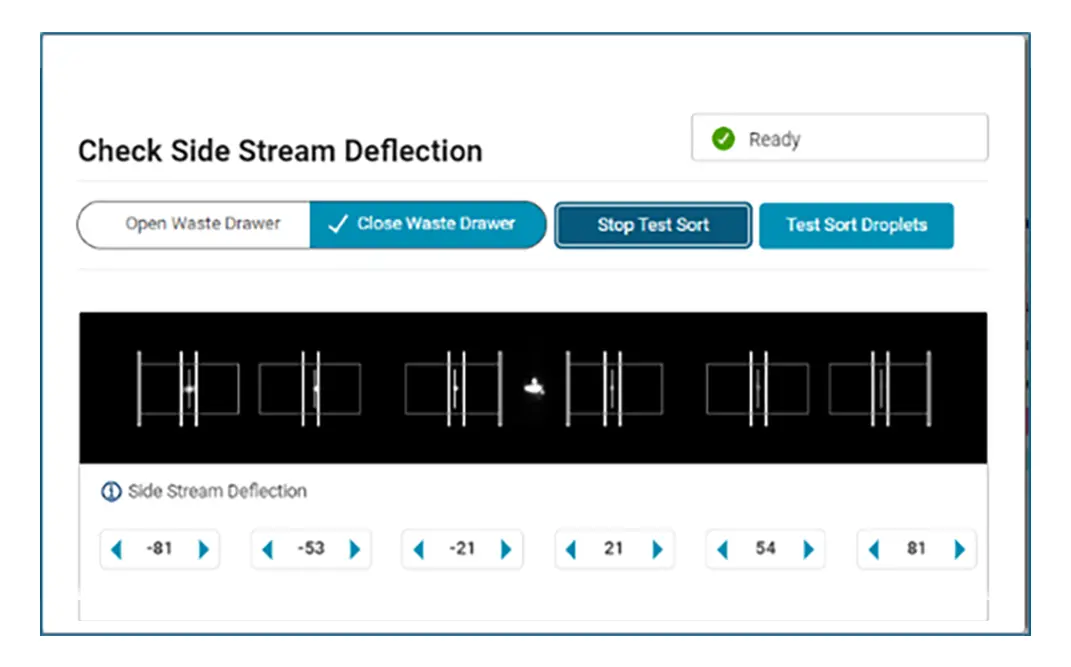
With automated stream setup, side stream tracking and BD FACSAccudrop™ Beads drop delay calculation in BD FACSChorus™ Software, more users can independently conduct their own sorts, giving them the opportunity to shine.
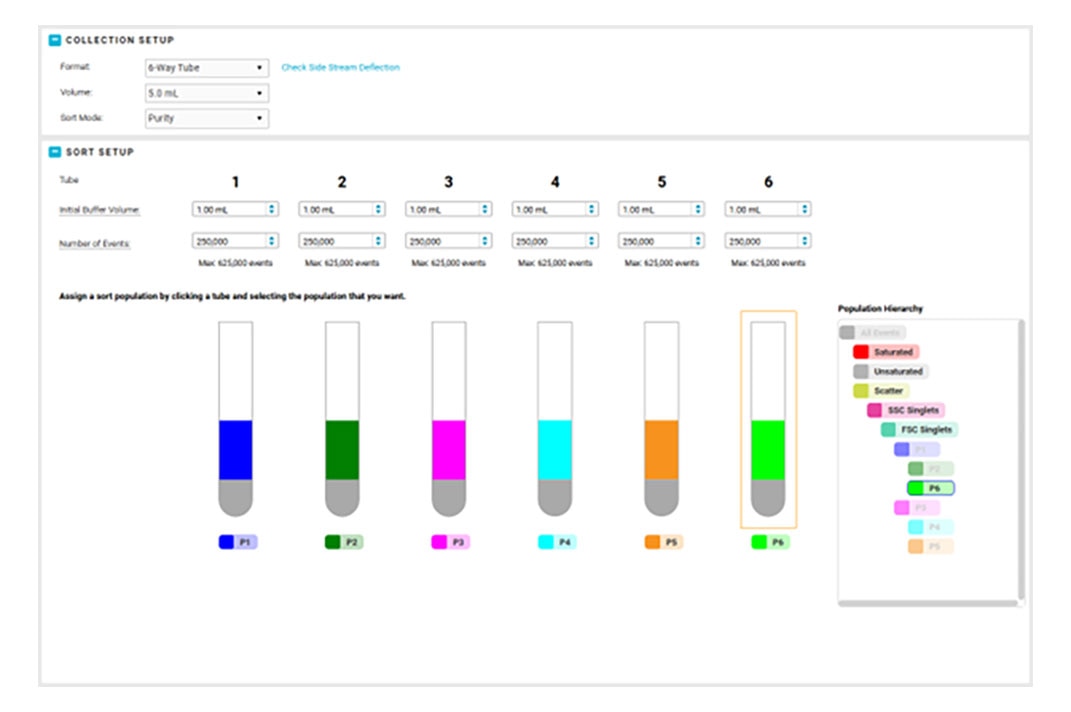
Combine that with the flexibility to adjust the side stream locations, sort into many sample collection types, modify the drop delay with an offset, and choose the nozzle size to meet your application needs, and BD FACSChorus™ Software helps your BD FACSDiscover™ S8 Cell Sorter ready to be the star performer your lab needs.
We know your lab is busy and your time is valuable. BD FACSChorus™ Software ensures you don’t waste time and gives you the gift of confidence as you acquire data on the BD FACSDiscover™ A8 Cell Analyzer.
First, with the built-in integrated autoloader, there are no manual hardware set up steps to enable acquisition from a plate or carrier. You can then choose either High-speed mode or Imaging mode for your experiments. Once your samples are loaded, you can step away from the system, allowing the automated system to safely detect and recover from any clogs or bubbles, ensuring your precious samples are handled with care.
Latest Features and Improvements
A smarter way to end your day: You can now end your loader runs on the BD FACSDiscover™ A8 Cell Analyzer with an automated walk-away cleaning and shut-down sequence. The system will run a routine clean, put the hardware into a safe standby state and log you out of BD FACSChorus™ Software v6.3.
Enhanced usability and workflow: This latest version is packed with improvements from start-up and connection to the experiment workflow and end-of-the-month usage tracking. Upgrade today to get the most from your BD FACSDiscover™ instruments!
New Sort Flexibility with BD FACSChorus™ Software
BD FACSChorus™ Software versions 6.2 and beyond support pausing a sort, unloading the sample tube and resuming without losing progress or resetting the sort counters. Additionally, you can add a drop delay offset to the calculated drop delay to sort large particles with more precision.
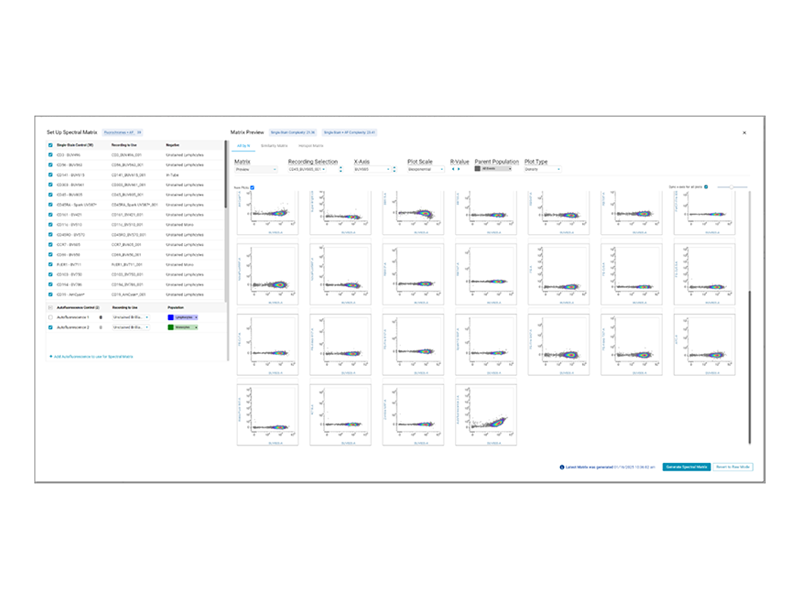
Introducing the New Spectral Unmixing Workflow with BD FACSChorus™ Software
BD FACSChorus™ Software improves the single-stain analysis and approval workflow, allows you to import single-stain controls from a previous experiment, plots the actual spectral signatures of your acquired controls, and enables you to create and manage multiple unmixing matrices for your experiment – all to ensure an efficient and accurate experiment run.
-
Flyer
-
White Paper
-
Abbreviation Chart
For Research Use Only. Not for use in diagnostic or therapeutic procedures.
Cy is a trademark of Global Life Sciences Solutions Germany GmbH or an affiliate doing business as Cytiva.
Refer to manufacturer's instructions for use and related User Manuals and Technical Data Sheets before using this product as described.
Comparisons, where applicable, are made against older BD technology, manual methods or are general performance claims. Comparisons are not made against non-BD technologies, unless otherwise noted.