-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD OMICS-Guard™ Sample Preservation Buffer
- BD® AbSeq Assay
- BD® OMICS-One Immune Profiler Protein Panel
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Accessory Kits
- BD® OMICS-One Protein Panels
- BD OMICS-One™ WTA Next Assay
- BD OMICS-Guard™ CRYO Preservation Buffer
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
-
Training
- Flow Cytometry Basic Training
-
Product-Based Training
- FACSAria Product Based Training
- FACSMelody Product-Based Training
- FACSLyric Product-Based Training
- FACSCanto Product-Based Training
- LSRFortessa Product-Based Training
- FACSymphony Product-Based Training
- FACSDuet Product-Based Training
- HTS Product-Based Training
- BD FACSDiscover™ S8 Cell Sorter Product Training
-
Advanced Training
Old Browser
This page has been recently translated and is available in French now.
Looks like you're visiting us from {countryName}.
Would you like to stay on the current location site or be switched to your location?
Redefine CITE-seq
The Intracellular CITE-seq Assay using BD® AbSeq Antibody-Oligos:
- Enables simultaneous RNA, surface protein and intracellular protein detection
- Supports high-plex proteomics profiling (~100 protein markers) at the single-cell level
- Allows a safe stopping point during the workflow without compromising assay sensitivity
- Works with the BD® Single-Cell Multiplexing Kit to enable sample multiplexing
- Drives deeper understanding of cellular functions such as signal transduction dynamics and transcriptional regulation
Get more information from the Intracellular CITE-seq Assay using BD® AbSeq Antibody-Oligos brochure.
List of Available Intracellular BD® AbSeq Ab-Oligo Specificities
| Specificity | Clone |
Caspase-3 | C92-605 |
PARP (Cleaved) | F21-852 |
H2AX (pS139) | N1-431 |
T-bet | O4-46 |
Granzyme B | GB11 |
Helios | 22F6 |
BCL-6 | K112-91 |
GATA3 | L50-823 |
Ki-67 | B56 |
Stat6 (pY641) | 18/P-Stat6 |
p38 MAPK (pT180/pY182) | 36/p38 (pT180/pY182) |
Chromogranin A | S21-537 |
Sox2 | O30-678 |
Sox17 | P7-969 |
Applications
Intracellular BD® AbSeq Antibody-Oligos can be used to characterize a heterogeneous subset of T follicular helper (TFH) cells in human tonsil
A combination of intracellular CITE-seq, surface CITE-seq and scRNA-seq was used to deepen our understanding of the differentiation states and subsets of cells present at low abundance, such as TFH cells.
We characterized the heterogeneous subsets of TFH and circulating TFH (cTFH) cells using a combination of intracellular CITE-seq, surface CITE-seq and whole transcriptome single-cell RNA sequencing (scRNA-seq) analysis. Germinal center and cTFH cells were enriched by sorting memory CD4+, CXCR5+, PD-1+ from a tonsil (n = 1) and memory CD4+ CXCR5+ cells from a matched blood sample and a healthy blood donor (n = 2). We examined the expression of Bcl-6 and other transcription factors, using protein and RNA expression data, across pre-TFH (an early development stage in the germinal center), TFH and cTFH cells.
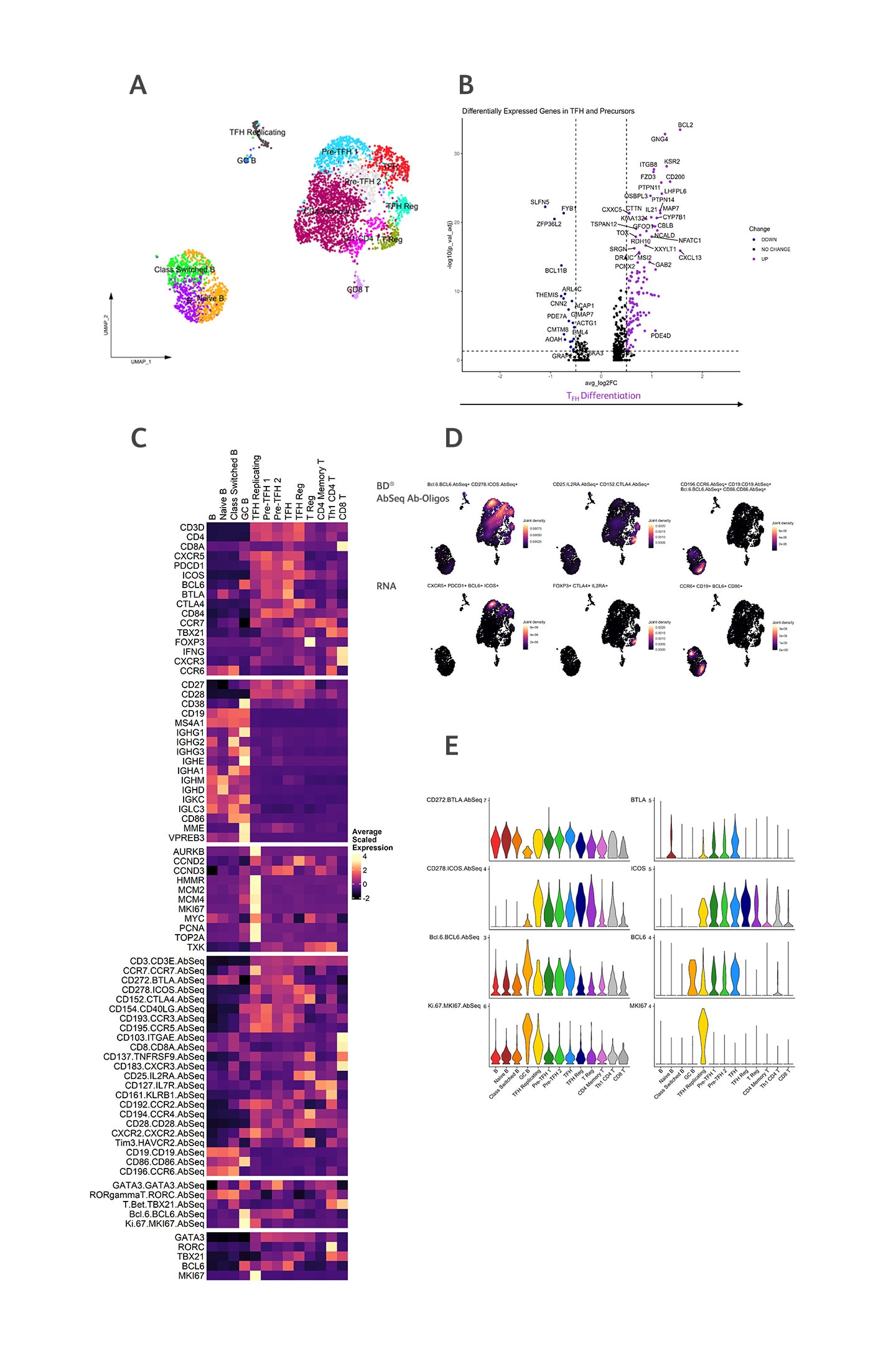
Tonsillar TFH cells were enriched and stained with both surface and intracellular BD® AbSeq Antibody-Oligos.
A) Annotated UMAP of tonsil, colored by cell type. B) Statistically significant (Bonferroni adjusted P value (P val adj) > 0.05) versus fold change for differentially expressed genes (DEG) between TFH cells compared to pre-TFH cells. C) Heatmap visualizing the gene signatures and surface and intracellular BD® AbSeq Antibodies used to identify cell subsets within the tonsil. D) Top panels display surface and intracellular BD® AbSeq Antibodies used to identify TFH cell (left), T regulatory cells (TReg) (middle) and B cells (right). Bottom panels display corresponding RNA signatures. E) Surface and intracellular BD® AbSeq Antibodies correlate with RNA expression within the tonsil.
