-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD OMICS-Guard™ Sample Preservation Buffer
- BD® AbSeq Assay
- BD® OMICS-One Immune Profiler Protein Panel
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Accessory Kits
- BD® OMICS-One Protein Panels
- BD OMICS-Guard™ CRYO Preservation Buffer
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
-
Thought Leadership
- News
- Blogs
-
Scientific Publications
-
Events
- Expanding PARADIGM to Infectious Disease Modeling: HIV & Tuberculosis
- CYTO 2023: Advancing the World of Cytometry
- Advances in Immune Monitoring Series
- Validating Flow Cytometry Assays for Cell Therapy
- Enhancing Cell Analysis with a New Set of Eyes
- BD Biosciences at International Clinical Cytometry Society 2025
Old Browser
This page has been recently translated and is available in French now.
Looks like you're visiting us from {countryName}.
Would you like to stay on the current location site or be switched to your location?
Uncover up to 30 immune markers in a single experiment
The BD® AbSeq Immune Discovery Panel (IDP) is developed using our antibody-oligo based technology and consists of 30 different specificities against most major human immune markers lyophilized in a single tube. The BD® AbSeq IDP is tested to detect human immune markers and is designed to work on the BD Rhapsody™ Single-Cell Analysis System with BD Rhapsody™ Single-Cell RNA Assays and BD® Multiplexing Kits.
Find more information from the BD® AbSeq Immune Discovery Panel brochure.

List of BD® AbSeq IDP Specificities
30 pre-titrated antibodies
| Specificity | Clone | Oligo ID |
|---|---|---|
| CD3 | UCHT1 | AHS0231 |
| CD4 | SK3 | AHS0032 |
| CD8 | SK1 | AHS0228 |
| CD11c | B-Ly6 | AHS0056 |
| CD14 | MPHIP9 | AHS0037 |
| CD16 | 3G8 | AHS0053 |
| CD19 | SJ25C1 | AHS0030 |
| CD25 | 2A3 | AHS0026 |
| CD27 | M-T271 | AHS0025 |
| CD28 | L293 | AHS0138 |
| Specificity | Clone | Oligo ID |
|---|---|---|
| CD45RA | HI100 | AHS0009 |
| CD56 | NCAM16 | AHS0019 |
| CD62L | DREG-56 | AHS0049 |
| CD127 | HIL-7R-M21 | AHS0028 |
| CD134 | ACT35 | AHS0013 |
| CD137 | 4B4-1 | AHS0003 |
| CD161 | HP-3G10 | AHS0205 |
| CD183 (CXCR3) | 1C6/CXCR3 | AHS0031 |
| CD185 (CXCR5) | RF8B2 | AHS0039 |
| CD186 (CXCR6) | 13B 1E5 | AHS0148 |
| Specificity | Clone | Oligo ID |
|---|---|---|
| CD196 (CCR6) | 11A9 | AHS0034 |
| CD197 (CCR7) | 2-L1-A | AHS0273 |
| CD272 | J168-540 | AHS0052 |
| CD278 | DX29 | AHS0012 |
| CD279 | EH12.1 | AHS0014 |
| CD357 (GITR) | V27-580 | AHS0104 |
| CD366 (TIM-3) | 7D3 | AHS0016 |
| HLA-DR | G46-6 | AHS0035 |
| IgD | IA6-2 | AHS0058 |
| IdM | G20-127 | AHS0198 |
BD® AbSeq IDP Protocol
Follow these simple steps to use the BD® AbSeq IDP:
Steps for reconstituting the lyophilized IDP and staining cells
- Remove the IDP tube from the foil bag and bring up to room temperature for 5 minutes
- Make sure the pellet is located at the bottom of the tube
- Add 35 µL of nuclease-free water to the bottom of the tube and allow antibodies to reconstitute for 5 minutes at room temperature
- Store the reconstituted antibodies on ice until the cells are ready for staining
Note: Reconstituted antibodies should be used within 24 hours. - To make 2X AbSeq labeling master mix, add 65 µL of BD Pharmingen™ Stain Buffer to the solution to bring it to a total of 100 µL
- Mix with 100 µL cell suspension prepared following Single Cell Labeling with BD® AbSeq Ab-Oligos (Doc ID 214394 Rev. 1.0) and stain the cells on ice for 30 minutes
- Add 3 mL BD Pharmingen™ Stain Buffer (FBS) to labeled cells and pipette-mix
- Centrifuge each tube at 400 x g for 5 minutes
- Uncap each tube and invert to decant supernatant into biohazardous waste. Keep the tube inverted and gently blot on a lint-free wipe to remove residual supernatant from tube rim
- Repeat steps 7–9 twice more for a total of three washes
- Resuspend pellet in 620 µL cold BD Pharmingen™ Stain Buffer (FBS) from the BD Rhapsody™ Cartridge Reagent Kit and proceed to single-cell capture
See the Single Cell Analysis Workflow with BD Rhapsody™ Systems.
PEFORMANCE
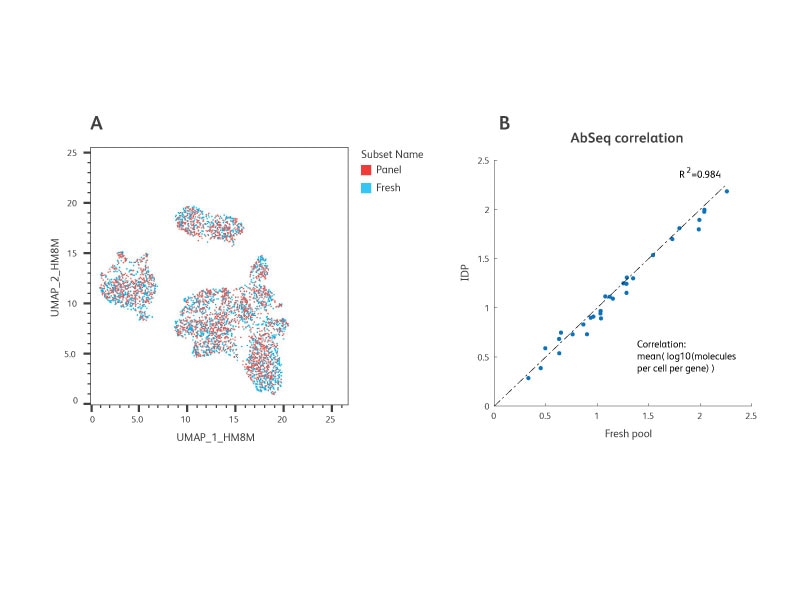
Similar performance between the IDP versus freshly mixed BD® AbSeq Antibodies. Following isolation from whole blood, peripheral blood mononuclear cells (PBMC) were split into resting (untreated) and activated (treated with CD3/CD28 for 24 hours) groups.
A 1:1 mixture of the resting and activated cells were stained with either the IDP or a freshly prepared mixture of the same AbSeq specificities. Equal number of cells were then loaded onto the BD Rhapsody™ Cartridges and AbSeq and WTA libraries were generated and sequenced (n = 2 individual experiments for this study). Data were analyzed using SeqGeq™ Software and Dataview Software.
A. UMAP demonstrated strong overlap in the cell groups identified between the IDP and fresh BD® AbSeq Antibody-stained samples.
B. The total number of AbSeq molecules detected by the IDP and fresh BD® AbSeq Antibody mixture showed a high correlation with R2 greater than 0.98.
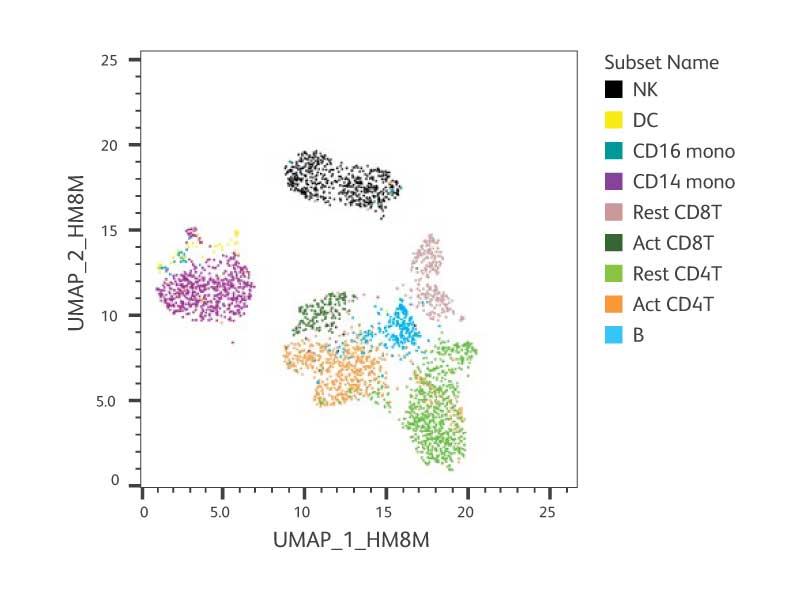
Performance of all 30 AbSeq specificities included in the IDP. PBMCs were activated and prepared as described in the figure above. After staining, cells were captured on the BD Rhapsody™ System and AbSeq and WTA libraries were generated and sequenced. To obtain over 80% sequencing saturation, the libraries were sequenced at 20,000 reads/cell for WTA and 30,000 reads/cell for AbSeq using the Illumina™ NextSeq™ High-Output Kit. Data were analyzed using SeqGeq™ Software and Dataview Software. We repeated the above experiments with at least two different donors. The representative figures from one donor are shown here.
A. Cell annotation of UMAP of resting + activated PBMCs resolved by the IDP antibodies and the WTA mRNA profile.
B. Heat maps of each AbSeq clone from IDP on UMAP from (A) showing the specificity of AbSeq detection for individual cell type.
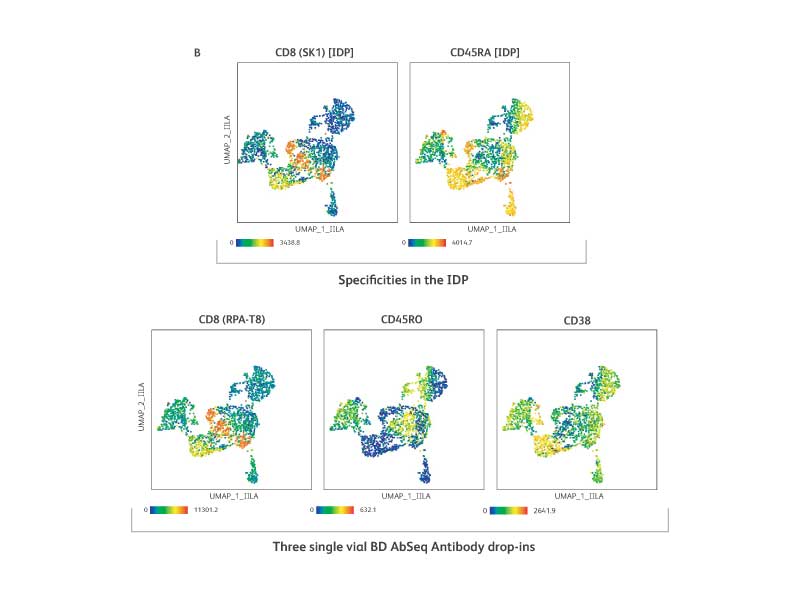
The IDP is a flexible backbone panel and accommodates additional AbSeq specificities. Three BD® AbSeq Antibodies were added and mixed with the reconstituted IDP pellet (n = 2).
A. The IDP performance was not impacted by drop-ins as shown by high correlation (R2 over 0.99 with or without drop-ins).
B. The IDP specificities of CD8[SK1] and CD45RA (top row) were assessed against the specificity of drop-ins CD8 [RPA-T8], CD45RO and CD38 (bottom row) and are shown in UMAP. Drop-in for CD38 detected cell types that are expected to be positive for CD38. Drop-in clone for CD8 (RPA-T8) showed a staining pattern very similar to the IDP clone (SK1) suggesting the high specificity of drop-in antibody as well as the compatibility of two clones for the same antigen. The contrasting expression pattern of CD45RO (drop-in) compared to CD45RA (IDP) further confirmed that adding the AbSeq specificities to the IDP had no adverse impact on experimental outcomes.
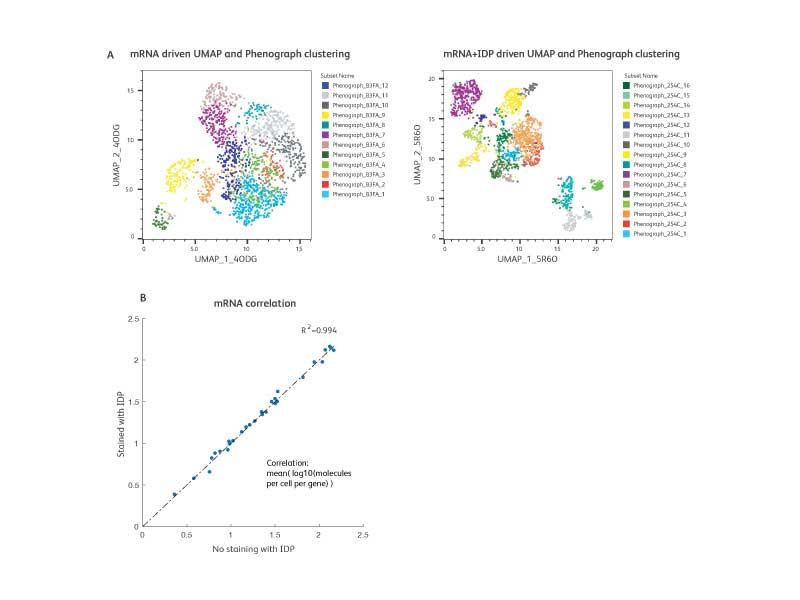
The IDP is designed to work with RNA and multiplexing assays
A. WTA and AbSeq libraries from IDP-stained cells (1:1 mixture of resting and activated PBMCs) were generated and sequenced. To illustrate the power of multiomic analysis, we analyzed the WTA data only (mRNA analysis) and compared with WTA + AbSeq data (mRNA and protein analysis). UMAP coordinates and unbiased clustering (Phenograph) using only WTA (mRNA) data are shown on the left, while coordinates and annotations using WTA + AbSeq (mRNA and protein) data are shown on the right. With a multiomics approach, additional cell types were revealed offering deeper biological insights.
B. To test the compatibility of the BD® Single-Cell Multiplexing Kit (SMK) and the IDP, we performed cell staining with the SMK and the IDP together and generated WTA, AbSeq and SMK libraries for sequencing. The expression of markers in the IDP was then compared to data generated in the absence of the SMK. These data showed that addition of the SMK does not impact the IDP as demonstrated by high correlation (R2 >0.99) between the IDP + WTA versus IDP + WTA + SMK.
-
Brochure
-
Data Sheets
-
A Comprehensive Characterization of Regulatory T Cells Using BD Rhapsody™ Single-Cell Analysis System
-
Assessing Obesity-Induced Inflammation in Mice through Single-Cell Multiomic Analysis
-
Exploring Tumor Heterogeneity of Chronic Lymphocytic Leukemia Using Single-Cell Multiomics
-
Simultaneous mRNA and Protein Quantification
-
Presentations
-
Protocols
-
Flow Cytometry Analysis of BD® AbSeq Ab-Oligo and Sample Tag Expression Protocol
-
mRNA Targeted and BD® AbSeq Library Preparation with the BD Rhapsody™ Targeted mRNA and AbSeq Amplification Kit Protocol
-
mRNA Targeted, Sample Tag, and BD® AbSeq Library Preparation with the BD Rhapsody™ Targeted mRNA and AbSeq Amplification and BD Single-Cell Multiplexing Kits Protocol
-
Preparing Single-Cell Suspensions Protocol
-
Single-Cell Capture and cDNA Synthesis with the BD Rhapsody™ Express Single-Cell Analysis System Protocol
-
Single-Cell Capture and cDNA Synthesis with the BD Rhapsody™ Single-Cell Analysis System Protocol
-
Single-Cell Labeling with the BD® Single-Cell Multiplexing Kit and BD® AbSeq Ab-Oligos Protocol
-
Single-Cell Labelling with BD® AbSeq Ab-Oligos Protocol
-
Single-Cell Labeling with BD® Single-Cell Multiplexing Kit and BD® AbSeq Ab-Oligos (41 plex to 100 plex)
To request a quote or place an order, email bdb_custom_orders@bd.com or contact your local BD sales representative.
For Research Use Only. Not for use in diagnostic or therapeutic procedures.
The information provided herein is not meant to be used, nor should it be used, to diagnose or treat any medical condition. All content, including text, graphics, images and information etc., contained in or available through this literature is for general information purposes only. For diagnosis or treatment of any medical condition, please consult your physician/doctor. Becton Dickinson India Private Limited and or its affiliates, its employees are not liable for any damages/claims to any person in any manner whatsoever.
