-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD OMICS-Guard™ Sample Preservation Buffer
- BD® AbSeq Assay
- BD® OMICS-One Immune Profiler Protein Panel
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Accessory Kits
- BD® OMICS-One Protein Panels
- BD OMICS-Guard™ CRYO Preservation Buffer
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
-
Thought Leadership
- News
- Blogs
-
Scientific Publications
-
Events
- Expanding PARADIGM to Infectious Disease Modeling: HIV & Tuberculosis
- CYTO 2023: Advancing the World of Cytometry
- Advances in Immune Monitoring Series
- Validating Flow Cytometry Assays for Cell Therapy
- Enhancing Cell Analysis with a New Set of Eyes
- BD Biosciences at International Clinical Cytometry Society 2025
Old Browser
This page has been recently translated and is available in French now.
Looks like you're visiting us from {countryName}.
Would you like to stay on the current location site or be switched to your location?
Request a Quote BD Rhapsody™ Single-Cell Analysis System
Please fill in the following information and we will get in touch with you regarding your query.
Overview
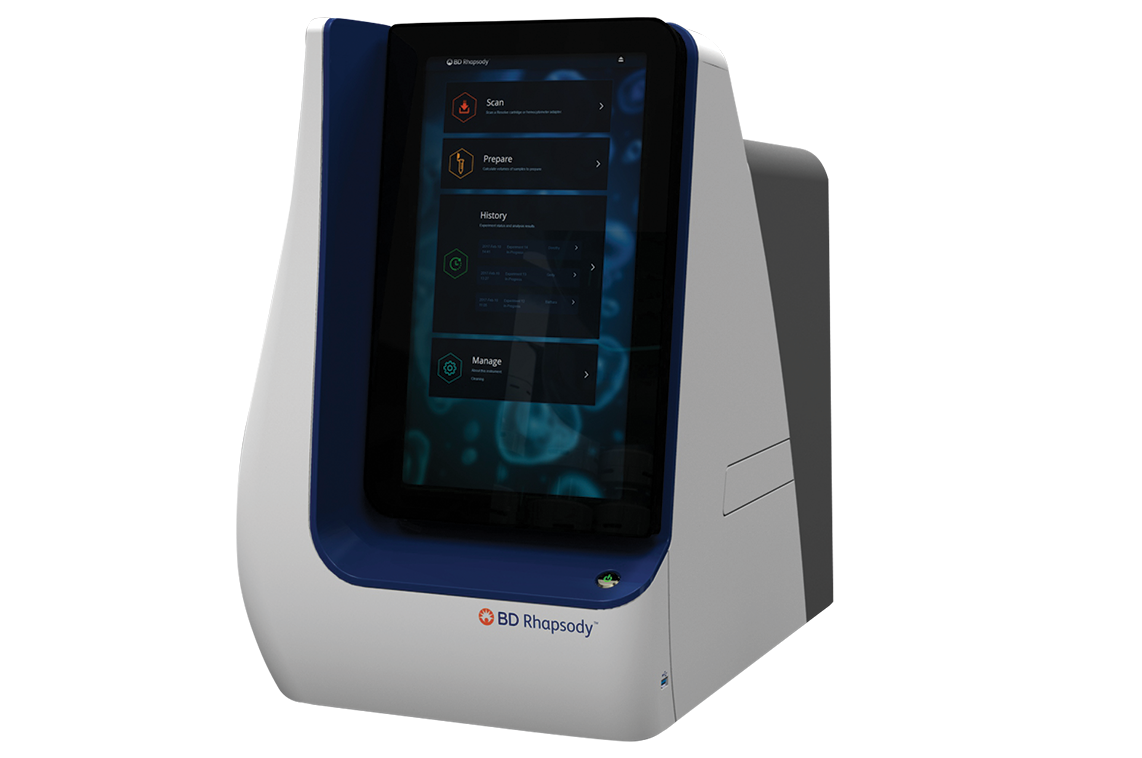
The BD Rhapsody™ Scanner is designed to visualize all steps in the single-cell capture workflow and provide detailed analysis metrics at every step, enabling the user to make key decisions throughout the workflow. The scanner has been redesigned to be compatible for use with the BD Rhapsody™ HT Xpress System and BD Rhapsody™ 8-Lane Cartridge for higher-throughput applications, enabling a multisample workflow. It can also be used in combination with the BD Rhapsody™ Express System and BD Rhapsody™ Single-Lane Cartridge for lower-throughput needs.
Get more information from the BD Rhapsody™ System brochure.
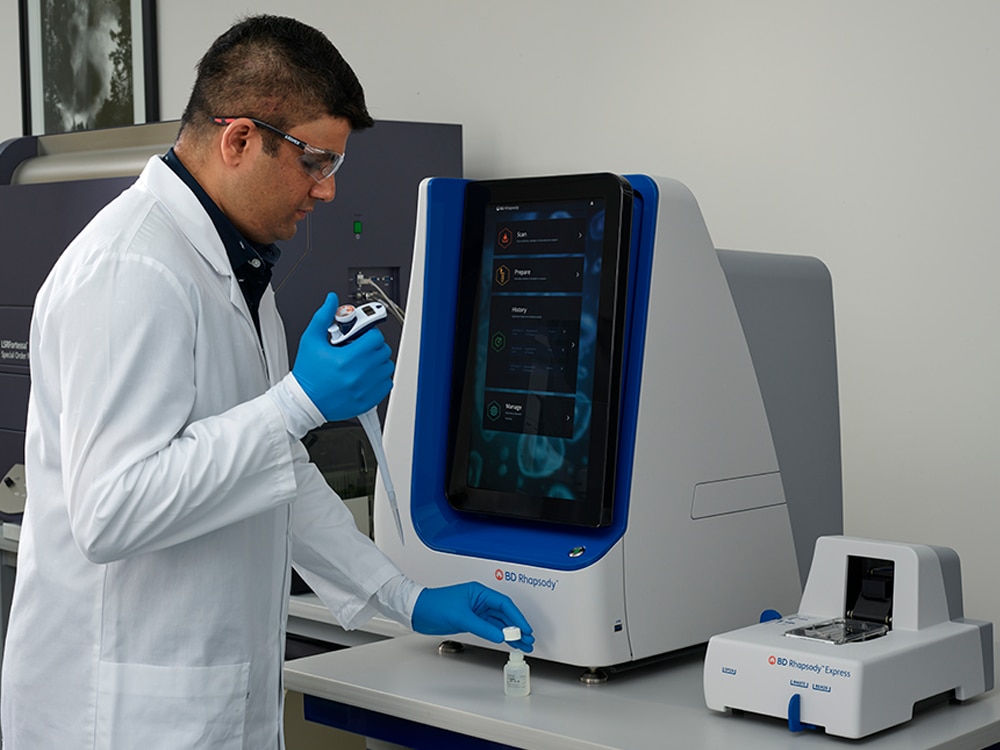

FEATURES

The BD Rhapsody™ Scanner provides the ability to visualize single-cell capture and provides key quality metrics at every step

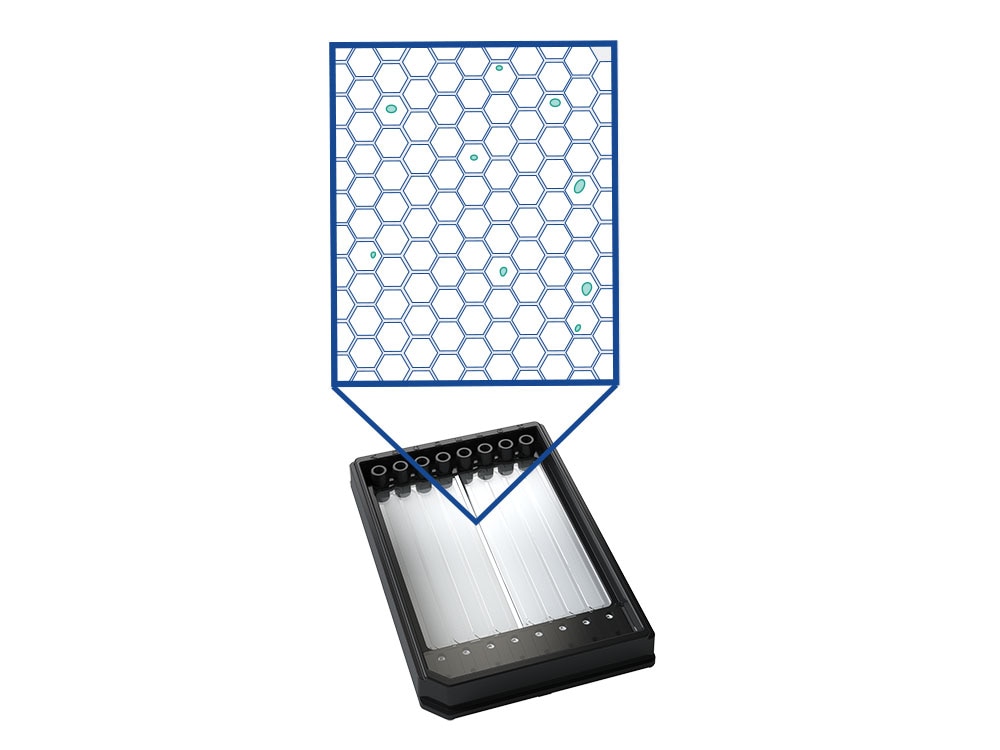
The BD Rhapsody™ Scanner enables visual inspection of the cartridge and of each microwell through an optically clear window on the BD Rhapsody™ Single-Lane or BD Rhapsody™ 8-Lane Cartridge
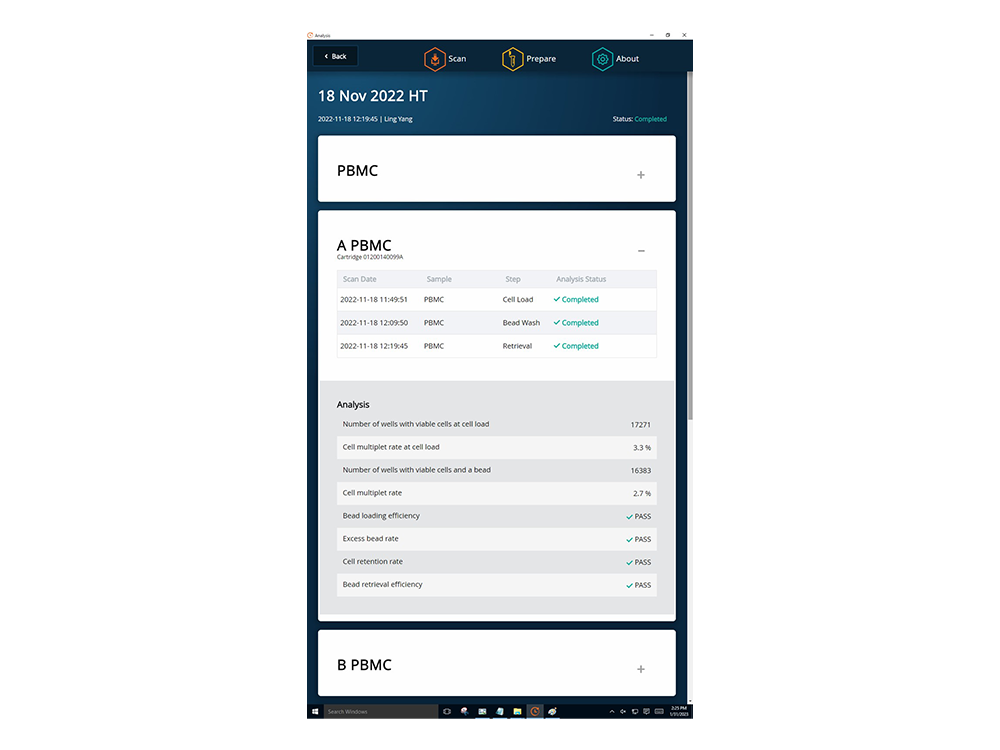
The BD Rhapsody™ Scanner provides the ability to visualize single-cell capture and provides key quality metrics at every step

The BD Rhapsody™ Scanner provides quality control measures such as cell capture rate and cell multiplet rates by direct imaging of the cartridge to analyze the quality of the cell preparation. Quality control metrics provide an indication if errors have occurred during the cartridge workflow allowing you to decide whether to proceed to sequencing.
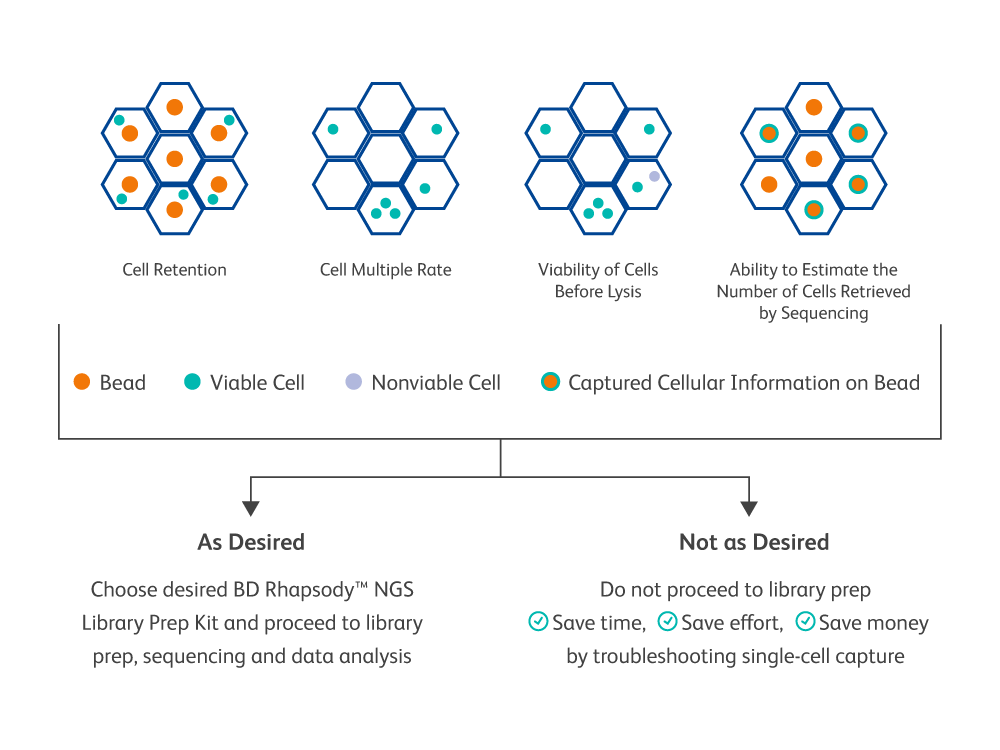
The BD Rhapsody™ Scanner provides the ability to visualize single-cell capture and provides key quality metrics at every step

The viability of the input sample and success of each step of the cartridge workflow can be confirmed, giving the user the power to decide whether to change course or troubleshoot, if necessary, before expensive downstream sequencing
BD Rhapsody™Single-Cell Analysis System Cartridge Workflow
BD Rhapsody™ HT Single-Cell Analysis System
-
Brochures
-
User Guides
-
Protocols
-
Technical Note
For Research Use Only. Not for use in diagnostic or therapeutic procedures.
Results shown are not guaranteed and may not be representative.
Dynabeads is a trademark of Thermo Fisher Scientific.
The information provided herein is not meant to be used, nor should it be used, to diagnose or treat any medical condition. All content, including text, graphics, images and information etc., contained in or available through this literature is for general information purposes only. For diagnosis or treatment of any medical condition, please consult your physician/doctor. Becton Dickinson India Private Limited and or its affiliates, its employees are not liable for any damages/claims to any person in any manner whatsoever.