-
Reagents
- Flow Cytometry Reagents
-
Western Blotting and Molecular Reagents
- Immunoassay Reagents
-
Single-Cell Multiomics Reagents
- BD OMICS-Guard™ Sample Preservation Buffer
- BD® AbSeq Assay
- BD® Single-Cell Multiplexing Kit
- BD Rhapsody™ ATAC-Seq Assays
- BD Rhapsody™ Whole Transcriptome Analysis (WTA) Amplification Kit
- BD Rhapsody™ TCR/BCR Next Multiomic Assays
- BD Rhapsody™ Targeted mRNA Kits
- BD Rhapsody™ Accessory Kits
- BD® OMICS-One Protein Panels
- BD OMICS-One™ WTA Next Assay
-
Functional Assays
-
Microscopy and Imaging Reagents
-
Cell Preparation and Separation Reagents
Old Browser
This page has been recently translated and is available in French now.
Looks like you're visiting us from {countryName}.
Would you like to stay on the current location site or be switched to your location?
The BD Rhapsody™ ATAC-Seq Assays enable you to either generate highly reproducible open chromatin profiling data from single cells or perform a multiomic analysis of open chromatin accessibility and transcriptome of single cells in one experiment using the BD Rhapsody™ Single-Cell Analysis System.
Are you interested in learning more about multiomic epigenomic profiling? Watch our webinar to learn how to combine image-based spectral cell sorting with ATAC-seq and whole transcriptome assays to decipher gene regulation in immunology research.
Learn more from the BD Rhapsody™ ATAC-Seq Assay Brochure.
Aymeric Silvin, a researcher from IGR (Institut Gustave Roussy) tested our kit and praised its ease of use and quick setup. He highlighted the high-quality data generated, making it an invaluable tool for researchers. Don't miss out on this revolutionary product - watch a short snapshot of the video.
Why Choose BD Rhapsody™ ATAC-Seq?
-
Unlock Chromatin Secrets: uncover a new layer of biology within chromatin accessibility patterns, gene regulation, and epigenetic dynamics - all on the same cell and with a wide range of cell inputs. -
Enhance cost Efficiency: the BD Rhapsody™ ATAC-Seq Assay has been designed to be cost-effective without compromising quality. -
Next-Level Insights: with our BD Rhapsody™ Single-Cell Multiomic ATAC-Seq Assay, you will reveal gene regulation networks with great sensitivity and specificity. -
Sample Tagging Enabled: compatibility with Custom BD® Nuclear Antibody-Oligonucleotide Conjugates allows to combine and simultaneously process multiple samples in one single reaction, further optimizing costs.
Features
Validated on the new BD Rhapsody™ Enhanced Cell Capture Beads V3, BD Rhapsody™ ATAC-Seq Assays generate consistent, high-quality data across different samples and users and a wide range of cell inputs. Key features include:
- High sensitivity and specificity across different experimental conditions
- Capable of accommodating a wide range of cell inputs
- Integrated epigenomic and transcriptomic characterization on the same cells
- Compatibility with Custom BD® Nuclear Antibody-Oligonucleotide Conjugates
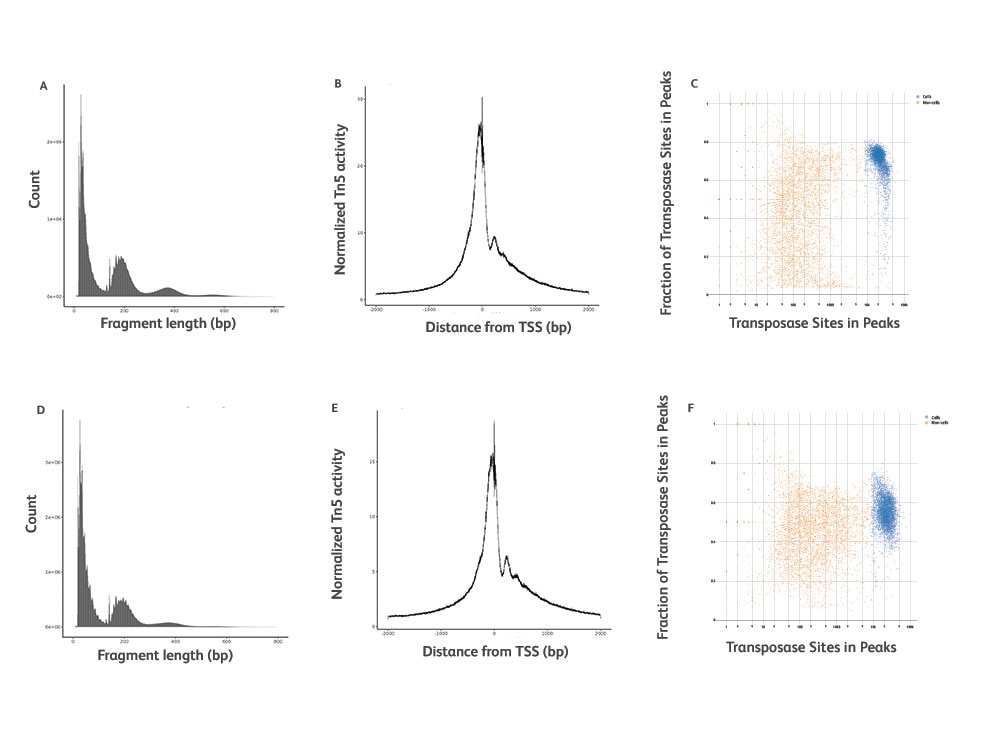
Unveil Epigenomic Heterogeneity with Great Precision

Sensitivity and specificity metrics for the BD Rhapsody™ Single-Cell ATAC-Seq Assay.
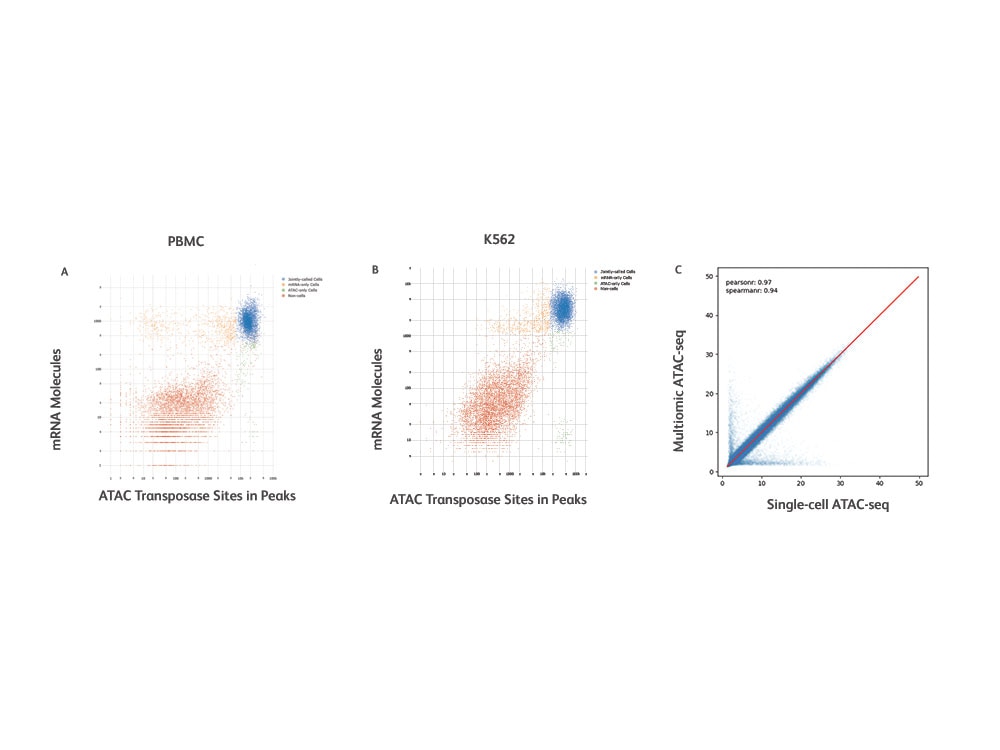
Uncover Ties Between Gene Regulation and Expression at the Single-cell Level

Sensitivity and specificity metrics for the BD Rhapsody™ Single-Cell Multiomic ATAC-Seq Assay.
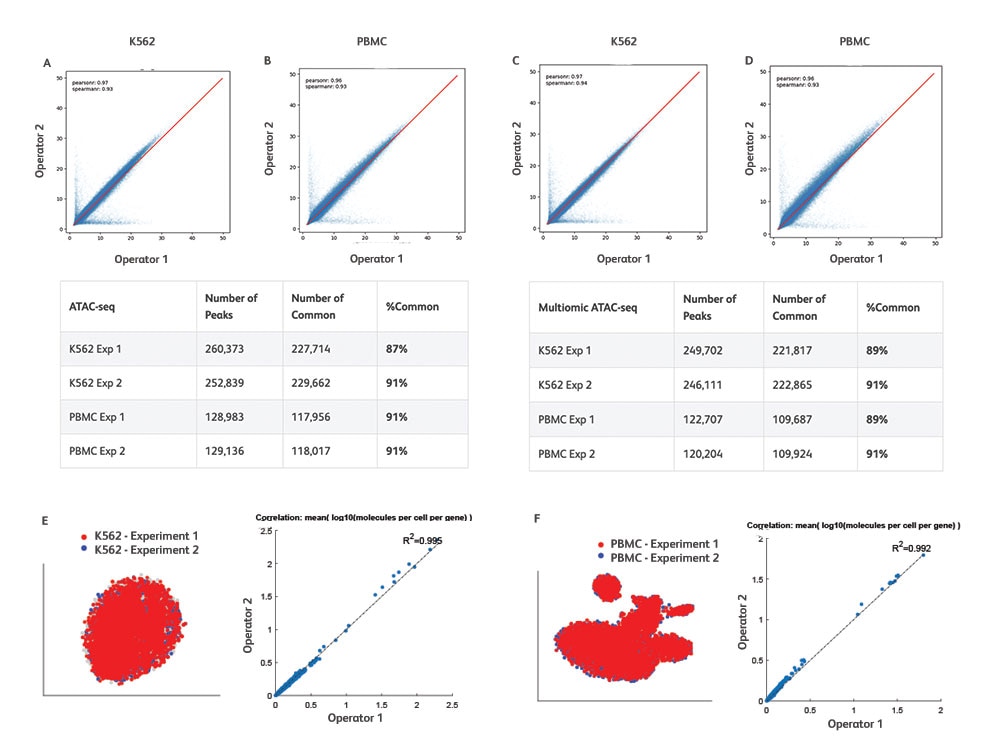
Reproducible Results Across Different Samples and Studies

High reproducibility with BD Rhapsody™ ATAC-Seq Assays
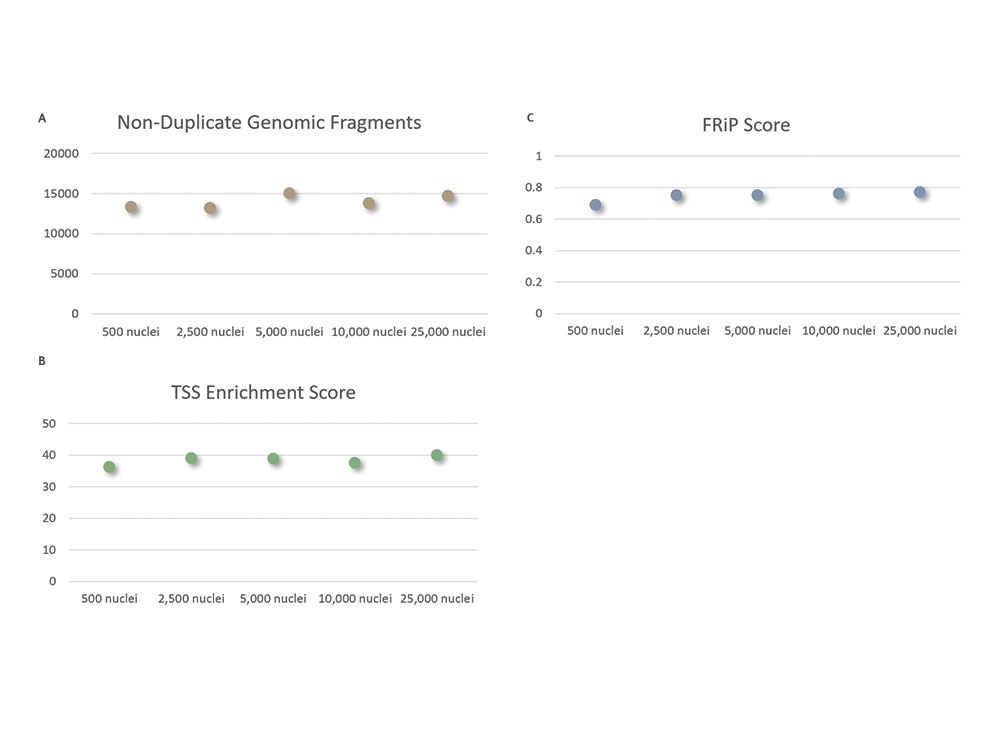
Generate Consistent, High-quality Data Across a Wide Range of Cell Inputs

Robust performance across different cell input amounts.
Applications
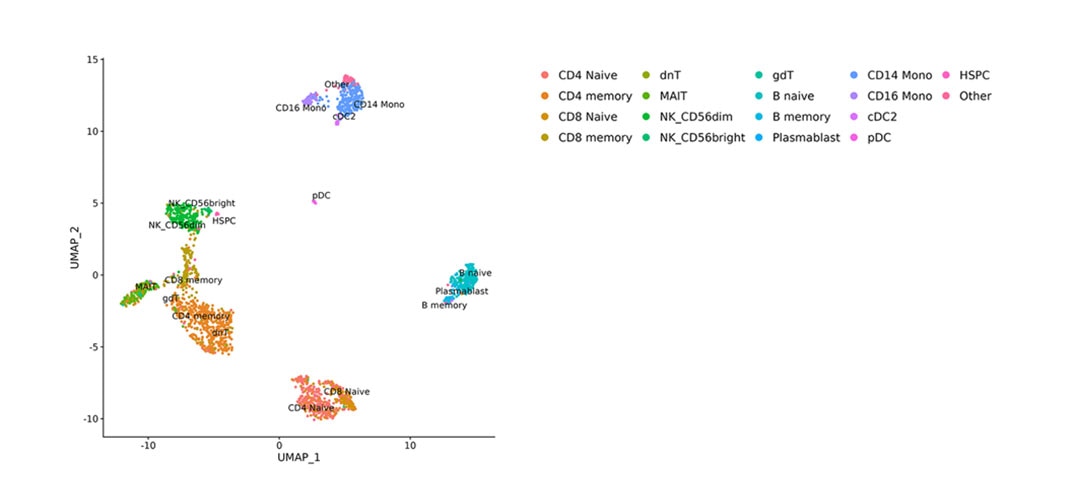
In this study, the BD Rhapsody™ Single-Cell Multiomic ATAC-Seq Assay was used to analyze 2,190 nuclei from human peripheral blood mononuclear cells (PBMCs). A joint whole transcriptome analysis (WTA) and ATAC-seq dimensionality reduction was performed by Uniform Manifold Approximation and Projection (UMAP) and cells were annotated as members of distinct cell types using the WTA data from a PBMC reference atlas.
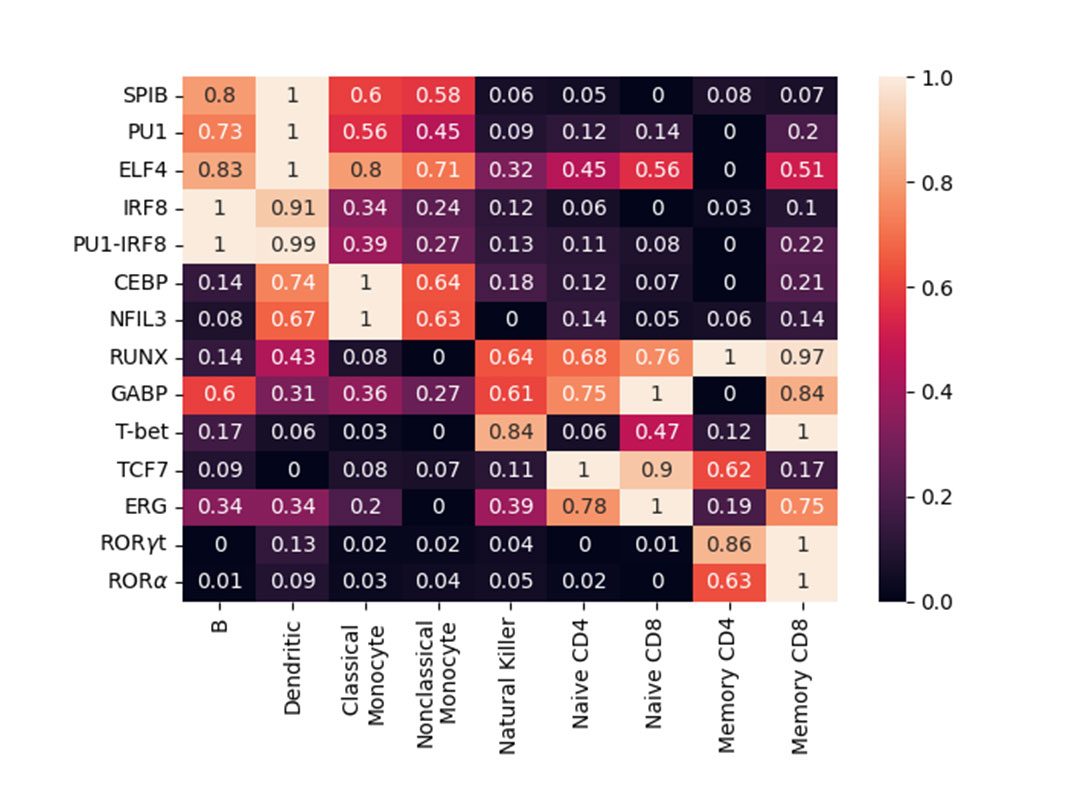
The heat map shows normalized enrichment scores of cell type–specific transcription factor motifs. Motif scores were calculated using a binomial distribution, determining the relative enrichment of each motif in differentially accessible regions of a given cell type compared to GC-matched background regions. The scores were then normalized across cell types per motif on a 0–1 scale, where 0 indicates least enrichment and 1 indicates highest enrichment of each motif.
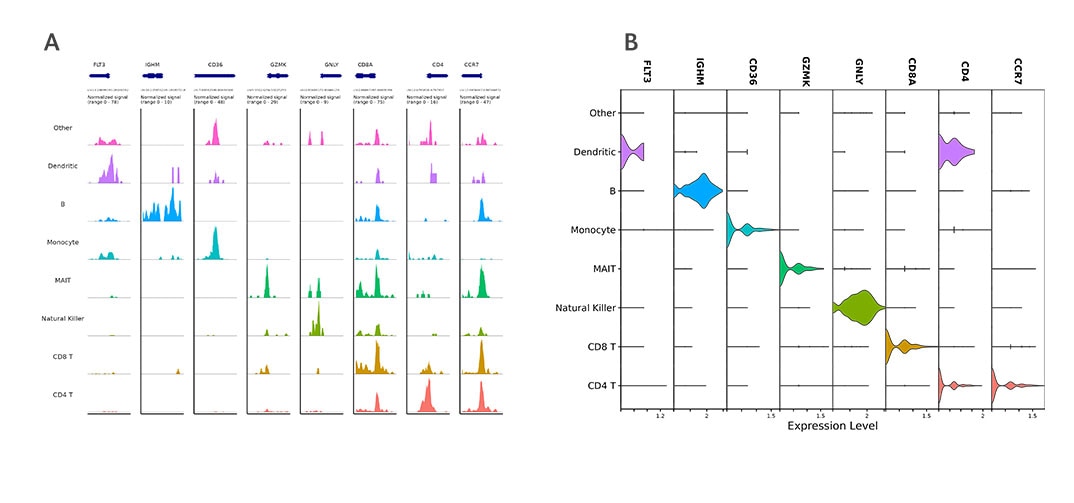
A. Read density across each ATAC-seq cluster at the transcription start sites of cell type marker genes.
B. Violin plots showing cell type-specific gene expression in WTA data.
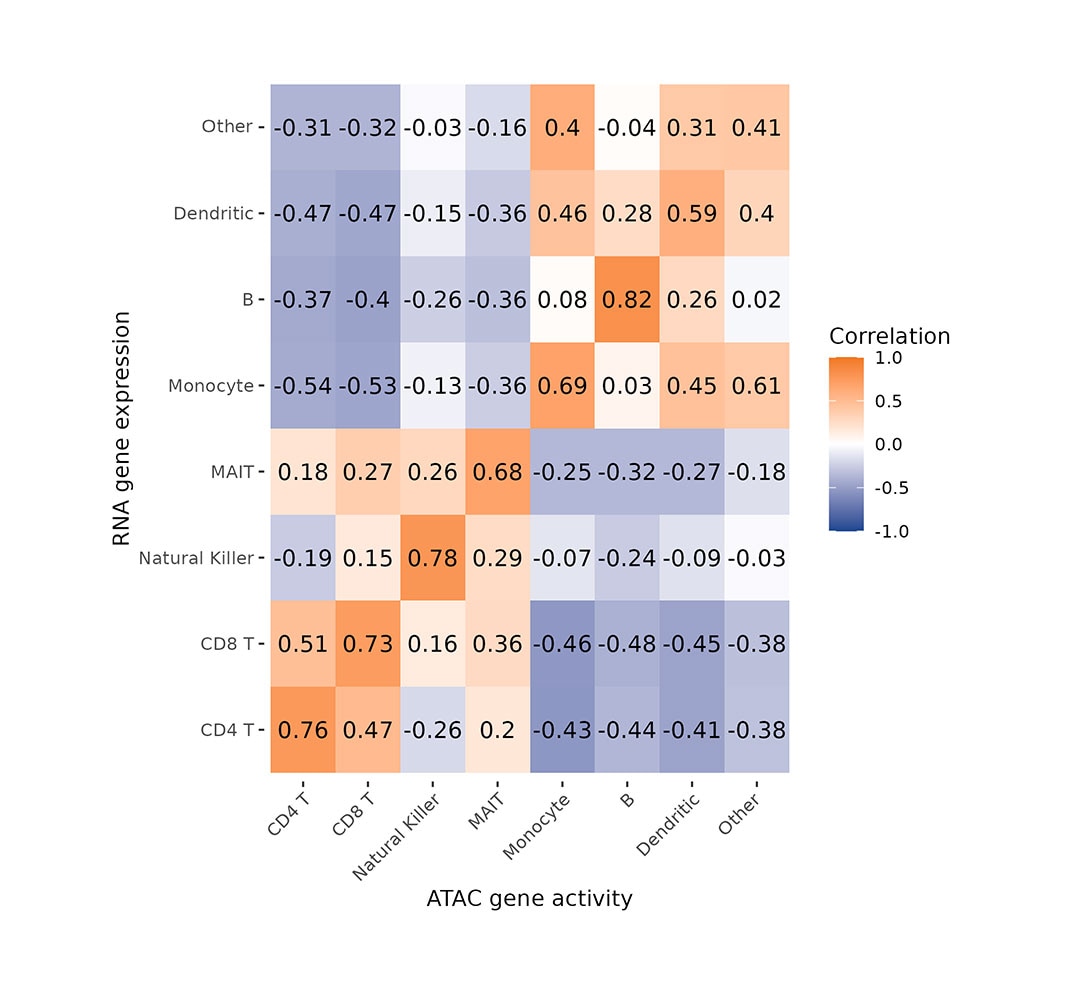
This heatmap shows Pearson’s correlation coefficients between ATAC-seq gene activity scores and gene expression values in PBMCs, with each row representing a cell type in WTA data and each column a cell type in ATAC-seq data.
For Research Use Only. Not for use in diagnostic or therapeutic procedures.
